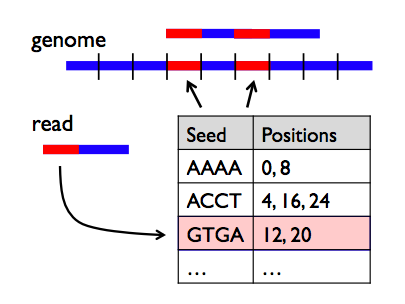
Why do we need Multiple sequence alignment (msa)? Local alignments are often preferable, but can be more difficult to calculate because of the additional challenge of identifying the regions of similarity. By contrast, local alignments identify regions of similarity within long sequences that are often widely divergent overall. Global alignment is a form of global optimization that "forces" the alignment to span the entire length of all query sequences. Those similar sequences that can emerge from different organism, often perform similar or even identical functions and at other times, mutating or rearranging to perform an altered function through the forces of natural selection. In a phylogenetic tree, each node with descendants represents the most recent common ancestor of the descendants, and edge lengths correspond to the numbers of changes between adjacent node in a tree.Ī sequence of amino acids which make up given segment of a protein molecule.Ī sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity that may be a consequence of functional, structural, or evolutionary relationships between the sequences. Both matrices estimate the rate at which each possible residue in a sequence changes to each other residue over time.Ī binary tree showing the represnting the relationship among a group of related nucleic acid or protien sequences from various/same species or other entities that are believed to have a common ancestor. 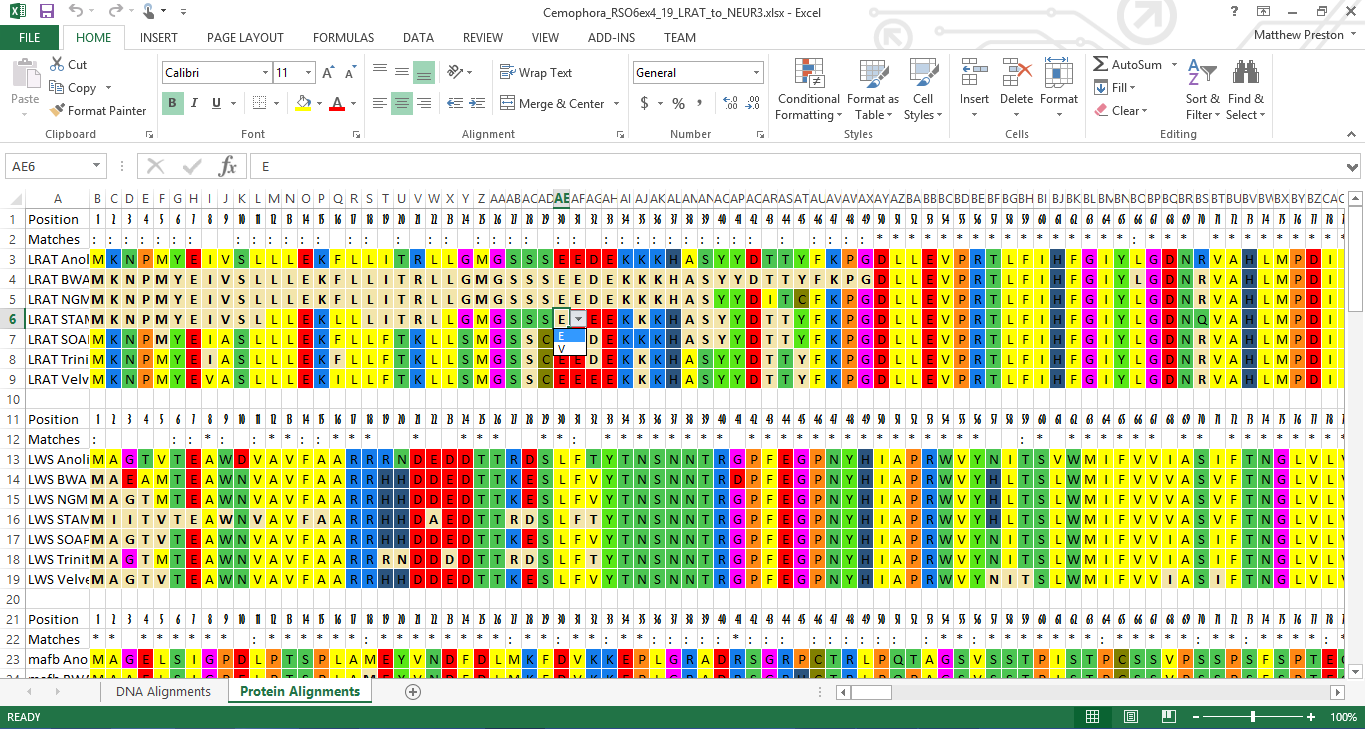
Substitution matrices used for sequence alignment of proteins. PAM (Point Accepted Mutation) / BLOSUM (BLOcks SUbstitution Matrix) Matrices A sequence of nucleotide bases in the DNA molecule.Ĭonserved pattern of amino acids that is found in one or more proteins, and
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
